ChemDoodle 3D
ChemDoodle 3D
3D Chemical Graphics, Animations and Modeling
for Windows, macOS and Linux
Click the play button to watch an introduction video

Looking to create 2D drawings for molecules and chemistry?
Check out ChemDoodle 2D →↓
Main Features
ChemDoodle 3D is a powerhouse for working with chemistry in 3D with industry leading molecular modeling tools and best-in-class graphics.
- 3D Visualization

A new perspective on chemistry
- Beautiful Scientific Models
We spend a very long time scrutinizing the models generated in ChemDoodle 3D. Take a close look at the renderings as you are rotating them to watch bonds orient towards the camera; every object mesh in the scene is built to merge with the others; and our models are some of the most gorgeously crafted by any software in this industry.
- Completely Customizable
All of the rendering in ChemDoodle 3D is controlled by styles defined by you. The styles can apply to the entire scene, selected content or individual objects.
- Advanced Text System
ChemDoodle 3D’s advanced 3D graphics engine can render text beautifully. Use this to show atom labels and more. You have full control over font, size and color.
- Custom Element Color Sets
In addition to the predefined Jmol, Rasmol, PyMOL and CDK color sets, you can also manually define your own elemntal color set to apply to molecule rendering.
- Compasses
Include a compass on the bottom left of the graphic, or in the center overlaying the content, to describe orientation.
- Projections
Both orthographic and perspective projections are available and can be switched instantly.
- Mesh Quality
You can quickly change between various quality levels to benefit graphics output or to speed up performance.
- Transparency
Transparency effects are just that, they let you see through objects in the scene. ChemDoodle 3D allows you to render transparent objects with or without back faces.
- Surfaces
ChemDoodle 3D can generate a number of surface types and color functions for any number of atoms. Visualize the space of your structures in ways other editors cannot.
You can customize mesh algorithms, display types, normals, colors and more.
Different types of surfaces are available, including van der Waals (VDW), solvent accessible surfaces (SAS) and solvent excluded surfaces (SES, Connolly).
Color your surfaces through various functions to provide a physical perspective on the structure. Functions include by atom color, charges (Gasteiger, QEq, QTPIE), lipophilicity (AlogP98) and molar refractivity (AMR98).
- Orbitals
View atomic orbital models based on quantum numbers with the Orbitals widget. Many options are available for customizing the graphics, which can then be output to an image. A great way to view the different orbital shapes.
- Gamma Correction
Gamma correction alters the brightness model of the graphics to compensate for the non-linear brightness physics of your monitor. ChemDoodle 3D defaults to a standard value of 2.2, but you can change this as appropriate for your screen.
- Art

Easily create ultra-realistic images, sketches, cartoons, paintings and more artistic illustrations
- Shader Programs
Shader programs are advanced instructions sent to the graphics card on how 3D graphics are generated. ChemDoodle 3D provides you with several built-in shaders to render your graphics, to create realistic styles like plastic and unrealistic styles like cartoons. Shaders in ChemDoodle 3D also provide additional features, such as outlining and shadows.
- Lighting
Control the color and direction of the lighting.
- Shading Model
A shading model is the method for converting the scene information into colors implemented by the shader program. Ambient-Diffuse-Specular (ADS) theory can be implemented in many different ways to create shaders that produce realistic graphics and non-realistic graphics. ChemDoodle 3D implements the following shading model types to choose from, all based on the ADS shading model:
- None
- Flat
- Cartoon
- Gouraud
- Phong
- Blinn-Phong
- Renderers
A renderer is the complete procedure in which we communicate with the graphics card using one or more shaders to produce a final image. In the simplest procedure, a single call is made to the graphics card where we pass in the scene data and produce an image with a single shader program. This is what we call a forward renderer.
The downside of a forward renderer, is each pixel is calculated in isolation, limiting the techniques we can achieve. We can instead chain several rendering passes through the graphics card together, using the output of one complete pass as input into another. This allows the software to produce some advanced effects.
In ChemDoodle 3D, you will find styles for 3 different types of renderers: (1) the forward renderer (2) the deferred renderer and (3) after effects.
- Anti-aliasing
Anti-aliasing smooths out the rough pixel edges in graphics (known as jaggies). ChemDoodle 3D supports hardware Multisample anti-aliasing (MSAA) for the forward renderer and both Fast Approximate anti-aliasing (FXAA) and Subpixel Morphological anti-aliasing (SMAA) for the deferred renderer.
- Shadows
Fully customizable, real-time, dynamic shadows add a major sense of realism to your graphics.
- 3D Textures
Textures apply a surface styling to the objects rendered in the scene. This is done by defining some pattern in three dimensions, similar to how you would see three dimensional patterns if you were to cut into a block of wood. The objects are then crafted out of this pattern to produce the graphics, allowing you to produce a number of interesting styles. Choose from granite textures, wood, stripes and more.
- Fogging
Define fogging using linear, exp1 and exp2 algorithms. Give incredible depth to your graphics or focus on specific sections with these features.
- Screen Space Ambient Occlusion
Screen space ambient occlusion (SSAO) simulates the occlusion of ambient light in corners and crevasses of 3D objects. If you look in the corner of your room, it will appear slighly darker than the centers of the walls. This technique adds a significant amount of realism to the image and greatly helps with perception of topography. This technique is fully customizable in ChemDoodle 3D.
- Outlining
Outline effects are created by finding significant changes in cached data values from pixel to pixel. There are two types of outline algorithms provided in ChemDoodle 3D: Trace and Artistic. You can control colors, thicknesses, sketchiness and more!
- Depth of Field
A depth of field effect focuses in on a specific portion of your scene, emulating images made with camera lenses. You can define the beginning and ending focus depths as well as the extent of the unfocused blur and dilate effects.
- After Effects
After effects are advanced image filters. ChemDoodle 3D includes the following after effects:
- Blur (Gaussian)
- Blur (Kuwahara)
- Blur (Median)
- Dilate
- Edge Detection
- Pixelize
- Posterize
- Sharpen
- Modeling

Interact with your structures in real-time with advanced physical simulations
- Real-time Optimization
ChemDoodle 3D's molecular modeling procedures can be run in real-time (using the Minimizer widget) so you can interact with molecules while you are building them or easily change between conformations. Our modeling engine is fast and efficient, allowing you to quickly generate relevant 3D coordinates for your built structures.
- The Optimization Function
Define the optimization function used to minimize energies from several built-in search directions including steepest descent, conjugate gradients, FIRE, and BFGS, with several line search options.
- Accurate Force Field Implementations
ChemDoodle 3D includes implementations of several published force fields. Our implementations are some of the most accurate and consistent in the industry.
- Universal Force Field
The Universal Force Field (UFF) is excellent for quickly building partial and complete chemical structures for demonstrations and images as it can handle the vast majority of the periodic table. A complete implementation is included in ChemDoodle 3D.
- Merck Molecular Force Field
The Merck Molecular Force Field (MMFF94) and its static variant (MMFF94s) are both accurately implemented in ChemDoodle 3D. Use this force field to generate experimentally accurate geometries for measurements and calculations.
- General Amber Force Field
The General Amber Force Field (GAFF) is compatible with the Amber Force Field for proteins and nucleic acids.
- VSEPR Force Field
The Valence Shell Electron Pair Repulsion (VSEPR) force field is a very specific Points-On-a-Sphere (POS) force field. The entire purpose of this force field is to produce ideal shapes for VSEPR theory. The VSEPR force field is perfect for students and instructors to use and demonstrate why certain atom centers, along with electron pairs, lead to certain shapes in molecular structures. Up to 8 connections on a center and multicenter structures are acceptable.
Because the purpose of the VSEPR force field is to generate shapes, energy calculations are irrelevant to the user, as they have no physical significance. When using the VSEPR force field, the Minimizer widget will instead display the current molecular geometry and VSEPR shape of the atomic center you are editing.
- Scope
Most small molecule force fields are optimized for describing individual discrete molecular structures. ChemDoodle 3D therefore will optimize molecular structures separately and individually as you are editing them. However, you may wish to optimize several discrete molecular structures at the same time and in relation to each other. To do this, you can change the optimization scope to optimize the entire scene.
Please note, while you are able to optimize the entire scene, most force fields are not parameterized for multiple discrete molecular structures, and you will be relying on the through-space forces defined in the force field (mainly van der Waals and electrostatic), if defined at all. While minimizations of several discrete structures using through-space forces may look correct, please understand the theory behind the force field you are using and that any force field not specifically developed or parameterized for intermolecular forces will not lead to experimentally accurate coordinates.
- Charge Models
Several charge models are implemented in ChemDoodle 3D: Gasteiger-Marsili PEOE, MMFF94, QEq and QTPIE. You can also use these charge models to color molecular surfaces.
- Atom Typing
Atom typing is an essential step in the correct application of a force field. You can see what atom types are defined to the atoms in ChemDoodle 3D for the specific force field you are using. You may even output the atom types for use in other applications.
- Force Rendering
Forces can be rendered to show how the molecule will change given the force field gradient calculations.
- Algorithm Transparency
In addition to showing you the atom typing and force vector calculations, ChemDoodle 3D will let you know when a structure is not compatible with a specific force field by providing descriptive errors and warnings. Use all of this information for reliability and referencing.
- Parallel Processing
Parallel processing allows you to use the power of your multi-core CPU to speed up computations. The force fields in ChemDoodle 3D are specially designed to run in parallel. This means that your structure, which would normally take x seconds to optimize, will now only take a fraction of x seconds to optimize. Enable parallel processing in ChemDoodle 3D when you want to handle the optimization of a large structure or system. Understanding how parallel processing works and is implemented in ChemDoodle 3D is very important. We have an entire section covering parallel processing in the ChemDoodle 3D user guide.
- Thorough Documentation
New to molecular modeling? No worries. We have an entire chapter in the ChemDoodle 3D user guide dedicated to molecular modeling and how it is implemented. You will be optimizing structures in no time.
- Symmetry

Easily analyze and visualize the symmetry of your structures
Symmetry Analysis
Use the Symmetry widget in ChemDoodle 3D to analyze point groups, framework groups, orders and vibrational representations for your loaded or built chemical structures and selections of atoms.
Visualization
ChemDoodle 3D can render symmetry components, including inversion centers, rotation axes, reflection planes, and rotoreflection components. These renderings may be included in your scene, where you may define the colors, transparency, extent and other features of the graphics.
Animation
Easily orient and animate the symmetry components of your system using the Symmetry widget. This is a very useful tool for students and a great way to gain insight into your molecular structures.
Orbits
ChemDoodle 3D will separate your atoms into symmetrically equivalent groups, called orbits. You may even color your structure by orbit, for another method to portray the symmetry of your system.
Symmetrization
ChemDoodle 3D includes a Symmetrize function to correct the atomic coordinates of the scene to conform precisely to the symmetry of the underlying point group. Symmetrized coordinates enhance both the accuracy and efficiency of molecular analysis and computation.
Symmetry features produced in collaboration with Professor Dean Johnston and symotter.org.
- Stereochemistry

Let ChemDoodle 3D do all the hard work for you
- Advanced Geometry Perception
ChemDoodle 3D will automatically detect and understand the 3D geometric features present in your chemical structures. Tetrahedral centers, alkenes and other isolated double bond systems, allenes and both odd and even numbered cumulenes, and atropisomers are all expertly handled.
- Input/Ouput
If you have or need files that contain stereochemical information, ChemDoodle 3D is the perfect tool for you. Stereochemical information in 3D chemical formats is simply stored in the 3D coordinates of the atoms. However, 0D/1D and 2D formats store stereochemical information in different ways. ChemDoodle 3D automatically handles stereochemical information in and out of popular chemical file formats, like MDL CTfiles, SMILES, InChI and more.
It may not be possible to automatically enforce all of the stereogenic features of a chemical structure, if the structure has multiple features present and is heavily embedded. ChemDoodle 3D will warn you of such cases, and you may always use the real-time modeling engine to easily create your desired stereoisomer.
- 0D/1D Support
Generate 3D coordinates with stereogenic features enforced from input SMILES, InChI and SLN strings. Output SMILES, InChI and SLN from 3D chemical structures, including stereogenic features.
It may not be possible to automatically enforce all of the stereogenic features of a chemical structure, if the structure has multiple features present and is heavily embedded. ChemDoodle 3D will warn you of such cases, and you may always use the real-time modeling engine to easily create your desired stereoisomer.
- 2D Support
Generate 3D coordinates with stereogenic features enforced from 2D drawings (such as from MDL MOLfiles), automatically interpreted using IUPAC stereochemistry rules for 2D structures.
It may not be possible to automatically enforce all of the stereogenic features of a chemical structure, if the structure has multiple features present and is heavily embedded. ChemDoodle 3D will warn you of such cases, and you may always use the real-time modeling engine to easily create your desired stereoisomer.
- Best-in-class CIP Algorithm
The CIP rules have long been the standard for describing configurations of stereochemical features in a molecule. While flawed, they have seen many revisions over the decades and were clarified by the work of Paulina Mata. These rules were adopted by IUPAC for naming standards and fully described in the Blue books. The most recent CIP rules from IUPAC were then algorithmically analyzed and standarized by Hanson et al. to remove any ambiguities and describe a completely consistent system for CIP assignments.
ChemDoodle 3D implements all 6 current CIP rules as well as auxilliary desciptors and mancude ring support. Stereochemical features in your structures will be assigned "R", "S", "E", "Z", "M" and "P" descriptors. The CIP algorithm in ChemDoodle 3D is validated against the test suite provided by Hanson et. al. and is 100% accurate in all 300 test cases provided.
Hanson, R. M.; Musacchio, S.; Mayfield, J. W.; Vainio, M. J.; Yerin, A.; Redkin, D. Algorithmic Analysis of Cahn−Ingold−Prelog Rules of Stereochemistry: Proposals for Revised Rules and a Guide for Machine Implementation. J. Chem. Inf. Model. 2018, 58, 1755-1765. - Pseudo-asymmetric Stereochemical Features
Pseudo-asymmetric stereochemical features in your structure are also resolved by our CIP engine, including meso centers. These will be assigned the lowercase "r", "s", "e", "z", "m" and "p" descriptors.
- Cis/Trans Double Bonds
The less systematic "cis" and "trans" stereochemistry descriptors can also be assigned for double bonds in unambiguous cases.
- Automatically Resolve Stereochemical Configurations
ChemDoodle 3D will assign all of the stereochemical configurations in your chemical structures with a single menu click.
- Force Stereochemical Configurations
If you do not want to manually build the desired stereoisomer of your structure, ChemDoodle 3D will do it for you, if necessary and possible. Just tell it which configuration to apply for a given stereochemical feature.
- Real Time Modeling
You may use the molecular modeling engine and the Minimizer widget to alter, invert and explore the stereoisomers of your structure in real time.
- Models & Printing

Several models have been exported from ChemDoodle 3D and printed using a 3D printer. These are great for gifts and celebrating your own scientific discoveries!
Export the objects you create for 3D printing and other CAD applications
- Model Files
Export your ChemDoodle 3D creations into many popular 3D model file types:
- 3D Manufacturing Format {.3mf}
- Additive Manufacturing File Format {.amf}
- COLLADA Digital Asset Exchange {.dae}
- Extensible 3D Graphics {.x3d}
- Polygon File Format {.ply}
- STereoLithography {.stl}
- Virtual Reality Modeling Language {.wrl}
- Wavefront Object File {.obj}
- Feature Support
ChemDoodle 3D supports as much of each format as possible, including meshes, normals, materials, transparency, colors and multicolors.
Note some formats are less capable. For instance, the STL format cannot store colors, so use a more capable format like the VRML formats to suit your application. - Options
Format specific options are also implemented, such as both binary and ASCII support for STL and PLY files and caching and compression for VRML files.
- Small Molecules

Build and visualize your molecules in 3D
- Intuitive Controls
ChemDoodle 3D’s building controls are made to clearly model the atoms and bonds they manage. Copious visual feedback is provided. There are also many options for customizing the building tools to your preference.
- Advanced Selections
Intuitive selection tools allow you to quickly trace, select and edit objects in 3D. For even more accuracy, use the Selector widget for a comprehensive organization of all the objects in the scene, allowing you to precisely investigate and select content.
- 7 Bond Types
6 physically relevant bond types are provided to create any type of chemical representation. No other program compares.
- Smart Bonds
ChemDoodle 3D renders multiple bonds that are able to orient themselves towards the camera for the most descriptive graphics.
- Optimize Zone
When building 3D structures, ChemDoodle 3D will automatically suggest the best place in three dimensions to add an atom connection. This helps you quickly build molecules. You can then simply turn on the minimizer to optimize physical coordinates. No other program has an optimize zone like ChemDoodle 3D’s!
- Balls and Sticks
You have full control over how atoms and bonds are rendered, and you can quickly choose from predefined representations, such as van der Waals spheres, ball and stick, stick, wireframe and line.
- Show Labels
ChemDoodle 3D’s advanced graphics engine can render text beautifully. Use this to show atom labels. You have full control over font, size and color.
- Add Hydrogens
Hydrogens will be added for you to your structures in appropriate 3D locations.
- Implicit Hydrogens
Implicit hydrogens are automatically tracked for your structures, but you can also override them as appropriate.
- Attributes
Define charges, radicals and isotopic mass values to atoms. These values are also properly read in and written to chemical file formats.
- Electron Pairs
Electron pairs and single electron (radical) objects are rendered on your structure and automatically placed for you (you may also manually place them). Use electron pairs in conjunction with the VSEPR force field to model VSEPR shapes.
- Aromatic Representations
Use toruses for aromatic circles or use resonance bonds that automatically orient to the center of rings.
- Bond Deduction
Automatically detect bonds in chemical files that do not contain bond data. Commonly, molecule data files do not include any bond information. Files including PDB, XYZ and output from quantum mechanics programs do not officially store bond data. Instead of having to draw bonds by hand, ChemDoodle 3D contains tools for deducing covalent bonds based on just the atom elements and 3D coordinates alone. ChemDoodle 3D can even assign bond orders to those bonds based on several algorithms, including "All Single", Antechamber and Auto-UFF.
- Stereochemistry
Perceive stereochemistry for 3D structures. CIP stereochemistry is determined for chiral centers, double bonds and allenes/cumulenes using our advanced and accurate CIP algorithms. Common cis/trans stereochemistry can also be determined for double bonds.
- 3D Alignment
Align molecular structures in 3D. Choose the mapping algorithm, whether to overlap the structures, or allow inversions.
- Optimize Structures
Quickly generate 3D coordinates for your structures using several algorithms.
- Measurements
Measure distances, angles and torsions in your structures.
- Cheminformatics Functions
Quickly access powerful functions to help modify your graphics: saturation, Kekulization, ring perception, distance geometry embedding, and more.
- Database Lookup
Search databases (PubChem, ChemSpider, ChemExper) for chemical structures and drag them right into your 3D scene using the MolGrabber widget.
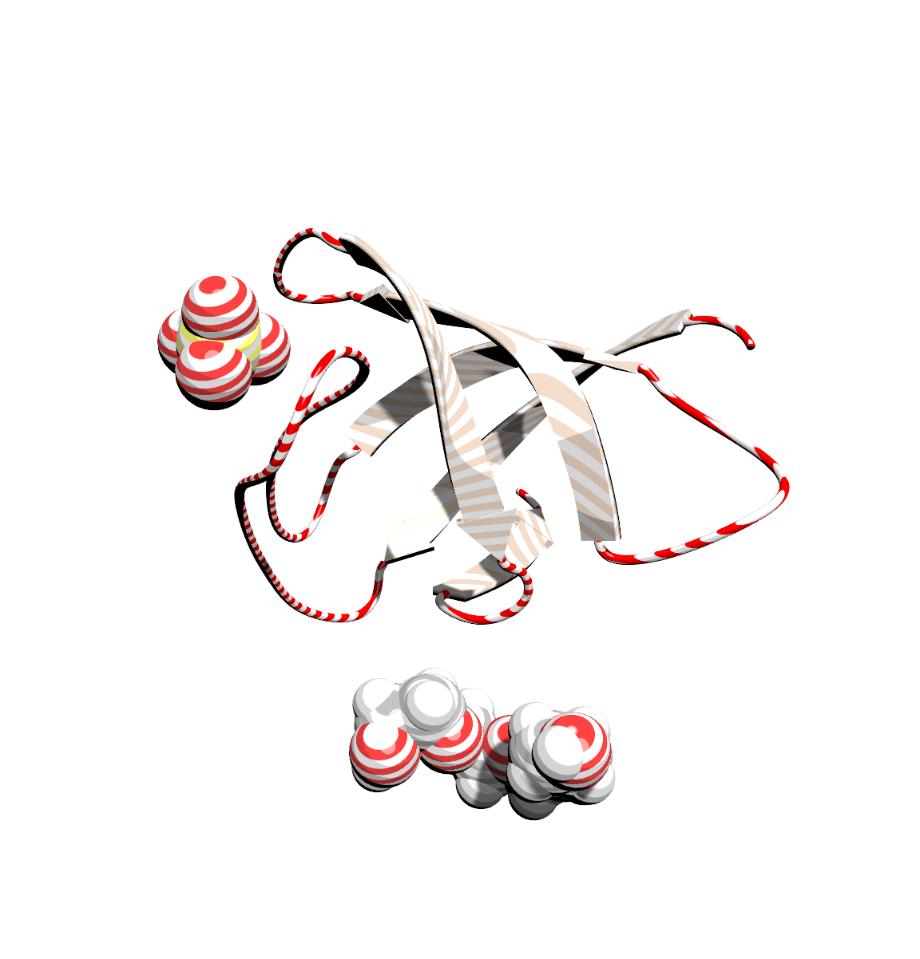
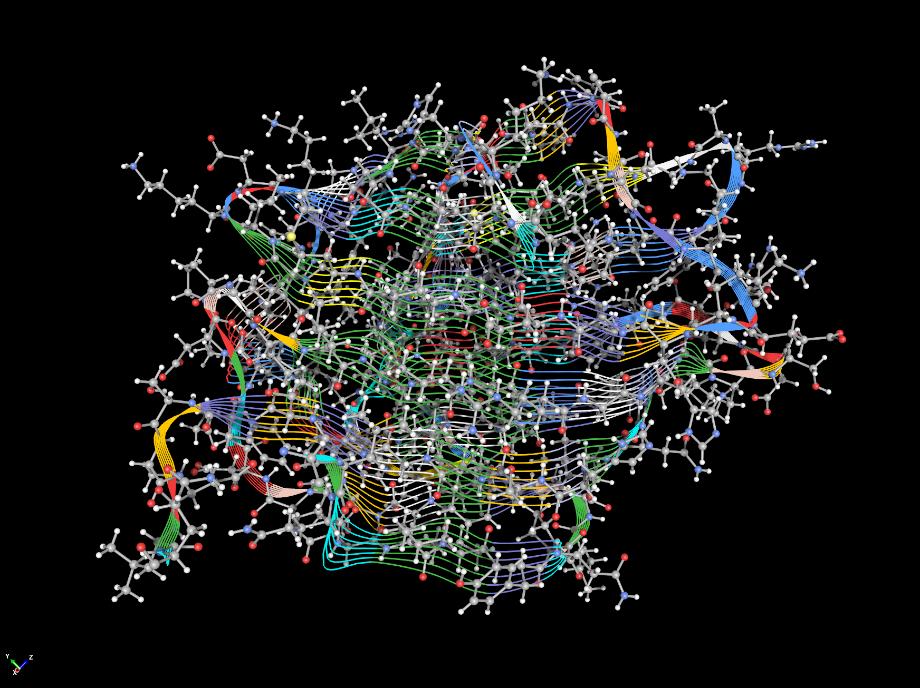
- Macromolecules

In addition to creating beautiful graphics for small molecule structures, ChemDoodle 3D will also help you to edit and create graphics for protein and nucleic acid macromolecules. The above image was rendered in ChemDoodle 3D of Protein Data Bank entry 5LRS with two solvent accessible surfaces rendered for the protein chains and using the None shader.
3D scientific graphics, big and small!
- Protein Data Bank Format
ChemDoodle 3D reads protein and nucleic acid information from PDB files and generates high quality meshes that are superimposed over the atom and bond coordinates.
- Macromolecular Transmission Format
ChemDoodle 3D also fully supports the RCSB Macromolecular Transmission Format for incredibly fast input of Protein Data Bank files with full bond order information.
- Protein Ribbon Models
Beautiful protein ribbons and traces are generated. Tubes, cylinder and plank models and cartoon models can also be generated. These models can be segmented, colored and completely controlled through styles. For instance, you can change the ribbon thickness or the alpha helix width.
- Nucleic Acid Ladder Models
Accurate nucleic acid ladder models are generated. These models can be colored and completely controlled through styles. For instance, you can change the backbone thickness or the platform height.
- Interpolation Algorithms
ChemDoodle 3D can use either a B-Spline algorithm or the Catmull-Rom algorithm for generating models. The Catmull-Rom spline creates more accurate models, but the B-Spline algorithm produces smoother and more aesthetic results.
- Residue Atoms and Bonds
Control residue atoms and bonds separately from the rest of the structures for unique graphics.
- Water
Show and hide water atoms defined in PDB files and represent them with stars instead of spheres.
- Periodic Systems

Unit cells are a great way to investigate repeating structures and crystals. The above image was rendered in ChemDoodle 3D of the zeolite Si-O framework, MFI, propagated along the z-axis with exponential fogging using the standard Blinn-Phong shader.
Infinitely repeating chemistry
- Crystallographic Information Format
ChemDoodle 3D reads periodic data from CIF files and can fully resolve point groups and symmetries. Unit cells for any symmetry geometry are extracted.
- Unit Cells and Extrapolated Supercells
Unit cells are rendered and supercells can be generated. Use an orthographic projection to get the best view.
- Nanotubes
Build armchair, zigzag and chiral nanotubes with several options. Periodic systems of nanotubes are automatically generated. A great way to output nanotube geometries for other applications.
- Stereoscopy

Augmented reality on your computer
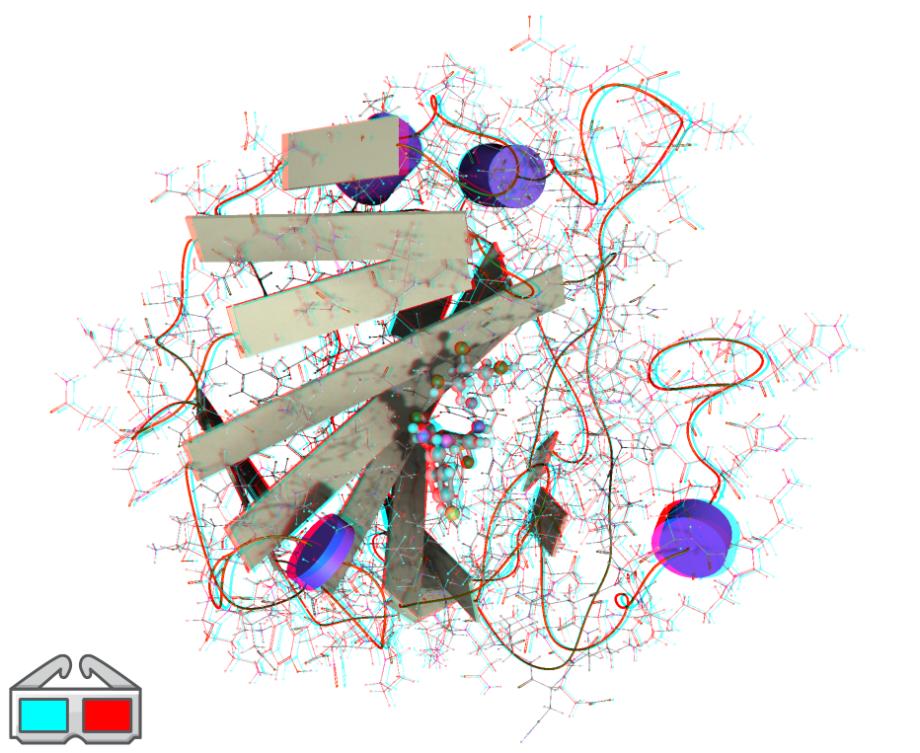
Anaglyphs
Anaglyphs are a stereoscopy technique used to show images in 3D space to our eyes from a 2D image. It works by rendering the scene for each eye's perspective in a different color space. Special glasses are then used to filter out each eye's image so one eye cannot see what the other eye sees. Our brain then pieces the two images together to observe the scene in 3D space.
Hardware
Anaglyphs are fully implemented with support for red-cyan, green-magenta, and amber-blue anaglyph glasses, which can be purchased cheaply from online retailers.
Customization
The Dubois method has been implemented for the most comfortable anaglyph viewing and you can control both the focal length and eye separation parameters to customize the anaglyph to your preference.
- Universality

ChemDoodle 3D is designed to perform everywhere for everyone
- Operating Systems
Fully supported on Windows, macOS and Linux.
- High DPI Support
Full support for high resolution displays, such as Retina Macs and the Microsoft Surface Pro.
- Light and Dark Mode Themes
Switch between interface "Light" and "Dark" mode themes to reduce strain on your eyes.
- Accessibility
Control interface colors, interface font sizes, define cursors and a cursor left-handed and right-handed mode make it comfortable for most users to use the software. You can also fully define all of the menu item accelerators (except the ones forced by the operating system) and tool shortcuts.
- Mobile Devices
A ChemDoodle license includes access to our ChemDoodle Mobile app companion for iOS (iPod/iPhone/iPad), Android and other mobile devices.
- 3rd Party Integration
ChemDoodle 3D works with several partners to help you improve your workflow. These partners include databases like PubChem, ChemExper and ChemSpider as well as ELNs such as LabArchives.
- Continuously Developed
We are always adding new features, and our customers continuously enjoy new updates with great new innovative solutions and tools that they have requested. Your license entitles you to all updates.
- Chemical Files
Read and write many popular 3D chemical file types for working with the applications you use:
Beilstein ROSDAL (.ros), Crystallographic Information Format (.cif), CHARMM CARD File (.crd), Chemical Markup Language (.cml), Daylight SMILES (.smi, .smiles), ChemDoodle 3D Scenes (.ic3), ChemDoodle Javascript Data (.cwc.js), HyperChem HIN (.hin), IUPAC InChI (.inchi), MDL MOLFiles, both V2000 and V3000 connection tables (.mol, .mdl), MDL SDFiles (.sdf, .sd), ORCA Input File (.orcainp, .inp), RCSB BinaryCIF (.bcif), RCSB Macromolecular Crystallographic Information File (.cif, .mcif, .mmcif), RCSB Protein Data Bank Files (.pdb, .ent), RCSB Protein Data Bank Markup Language (.xml, .pdbml), Schrödinger MacroModel (.mmd, .mmod), Tripos Mol2 (.mol2, .ml2, .sy2), Tripos Sybyl Line Notation (.sln), XYZ Files (.xyz)
- Bitmap (Raster, Pixel) Images
Write a large number of bitmap (also known as raster or pixel) images for use with other applications. Some formats have additional options, such as controlling output resolution in PNG, JPEG, and TIFF images.
- CompuServ Graphics Interchange Format {.gif}
- Joint Photographic Experts Group {.jpg, .jpeg, .jpe, .jfif, .jfi, .jif}
- Microsoft Bitmap {.bmp, .dib}
- Portable Network Graphics {.png}
- Tagged Image File Format {.tiff, .tif}
- UNIX Portable PixMap {.ppm, pnm, pbm, pgm}
- Wireless Bitmap {.wbmp}
- Web Components
Produce 3D ChemDoodle Web Components, which are high-quality, interactive, HTML5 components for websites and web apps that work in Microsoft Edge, Mozilla Firefox, Google Chrome, Apple Safari and Opera, and even on Mobile Safari on iOS and Chrome for Android.
- References
All of the resources we use to develop the algorithms in ChemDoodle 3D or the choices we make for the software are documented in the Help menu. This way you can evaluate the quality of our work.
- Thorough Documentation
A thorough user manual is free to download. It is in PDF format, fully searchable with a fully linked table of contents and index.
Screenshots
Try ChemDoodle 3D before you buy
A free trial for ChemDoodle 3D lasts for 14 days with some features restricted. The trial is not an obligation and we require no identifying or financial information to start a trial. After 14 days, the trial will no longer open. Our free trials are only for evaluation purposes and any output from the trial will contain a watermark and is copyright of iChemLabs.
To start your trial, do not buy, simply download ChemDoodle 3D for your operating system below, install and open it. Then accept the EULA. The next page will ask for a license code, instead click the I have not yet purchased a license code button and then the Free Trial button to begin.
Download v7.7.0
System Requirements
- Windows 10/11+, macOS 11+ (Big Sur, Monterey, Ventura, Sonoma, Sequoia, Tahoe or more recent), or Linux.
- A 64 bit (x86_64 or Apple Silicon) processor is required.
- ChemDoodle 3D requires OpenGL v2.1 drivers.
- A minimum of 1GB of memory, 2GB recommended.
Download User Guide